Daniel Polanco

Academically trained as a computational biologist. Open source contributor, and certified Apple App Store publisher. My long-term goal is to mentor and lead a software development team. In the near-term, to support and collaborate with clinical and bioinformatic research teams.
Work Experiences
Founder / Mobile Software Engineer
Collaborated with the Colorado Center for Personalized Medicine to redesign iMTracker (lifestyle tracking app) on iOS for a pilot study.
- Modernize, streamline, and improve perforamce of legacy user interface.
- General debug and software management, including unit testing, source control, and documentation.
- Modification of existing code to allow customization of notifications and direct data entry.
- Data storage organization to improve analysis performance and privacy control.
- Automate source build, code coverage, reporting, and app submission for publication.
Inovation Engineer
Created a holographic mobile app for HoloLens with Git, Visual Studio Team Services, and Agile methodology. Built and deployed on Azure.
Research Assistant
Developed and prototyped a model for identifying genomic duplications with Dr. David D. Pollock.
- Conduct statistical analysis and interpret expression levels from cDNA/mRNA.
- Locate and remove mitochondrial DNA, identify repeat sequences, and search for sequencing contamination.
- Concentrate, isolate, and clone plasmid DNA and further prepare DNA to be sent for off-site sequencing.
Projects
iMTracker
Design, engineer, and ready to publish a health personalization application that tracks symptoms. Symptoms are processed on devide (EDGE) and results are presented to summarize backend analysis.
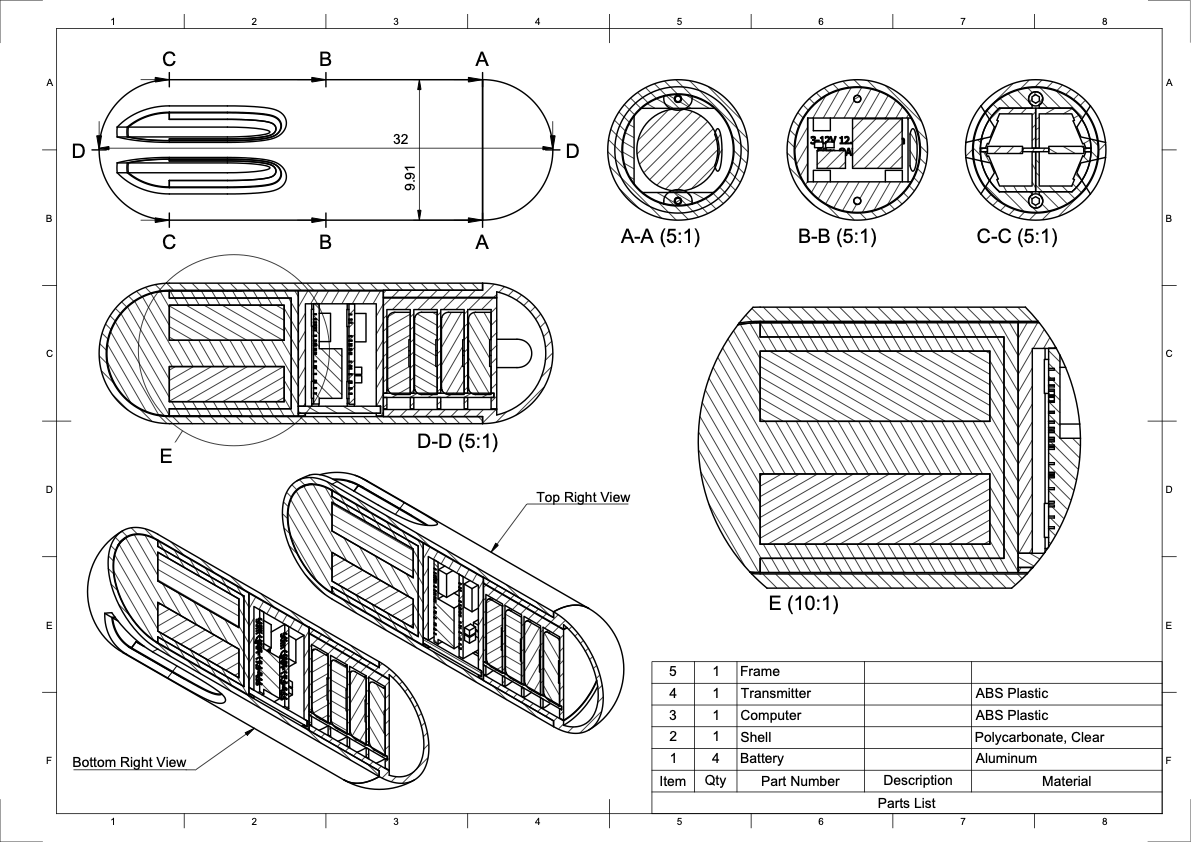
Enteroscan
Created a technical design for an ingestible pill including video demonstration. Created in Autodesk Fusion 360 for graduate coursework.
Duplication Identification
Identify genomic duplications using a novel algorithm for sorting and detecting key fragments.